Disambiguation
Link to this page as: http://alexmclark.com

 Alex Michael Clark, born in New Zealand, doctorate in Chemistry from the University of Auckland, lives in Ottawa (Canada).
Alex Michael Clark, born in New Zealand, doctorate in Chemistry from the University of Auckland, lives in Ottawa (Canada).
It's a common name, so if these descriptors don't match, you're probably looking for someone else.
Links
Primary blog: Cheminformatics 2.0 ⧉
ORCID: 0000-0002-3395-4666 ⧉
Social Media: LinkedIn ⧉, Twitter (X) ⧉
 Research Scientist at Collaborative Drug Discovery ⧉.
My role involves prototyping exotic new technologies and taking them through the lifecycle from idea to shipping product. These technologies
include molecular informatics, macromolecules, mixtures and bioassay protocols, among other things.
Research Scientist at Collaborative Drug Discovery ⧉.
My role involves prototyping exotic new technologies and taking them through the lifecycle from idea to shipping product. These technologies
include molecular informatics, macromolecules, mixtures and bioassay protocols, among other things.

Orangometallics
Demo link ⧉: a searchable catalog of inorganic molecules from vendor catalogs. These are uniquely drawn using correct valence rules and reasonable presentation aesthetics.COVID19 Models
Demo link ⧉: online models based on early COVID data, put together during the lockdowns of 2020.MedChem Toolkit
Demo link ⧉: a collection of utilities for medicinal chemistry, including structure prediction tools.Coordination InChI
Demo link ⧉: a working demonstration of algorithms that can create a unique string code for inorganic complexes, which serves as a proof of concept for adding this functionality to the InChI identifier.GoCatalysis
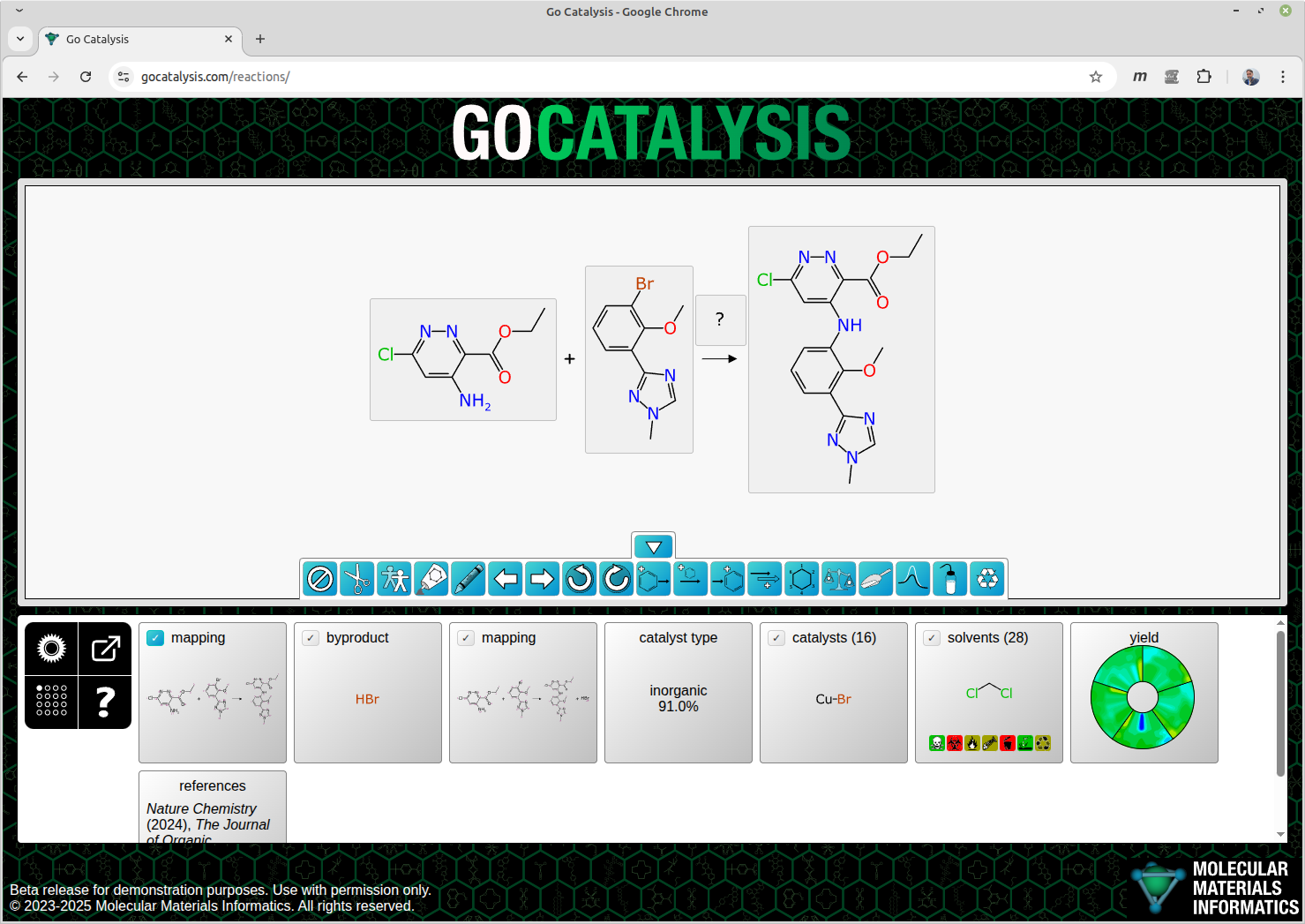
A prototype tool for predicting reaction conditions, such as ideal catalysts and solvents. More information is available in a series of blog posts ⧉. The service is current behind password protection, because the internet is a dangerous place, but anyone is welcome to take a look. Contact me by any means, such as email.
Open Source Software
GitHub accounts repository list ⧉
Open Source:
- WebMolKit ⧉: a general purpose cheminformatics toolkit written in TypeScript and available for all types of web-based applications and software built on the JavaScript runtime engine.
- Coordination InChI ⧉: proof of concept and validation data resources for InChI identifier extension to include inorganics.
- Mixtures ⧉: data container and tools for describing mixtures of chemicals.
- BioAssay Express ⧉: an ontology-based annotation tool for bioassay protocols.
Bibliography
A (mostly) complete list of peer-reviewed publications. The definition of peer review includes journal articles, book chapters, graduate theses, and patents, all of which have their chance to be rejected if peers deem them to be unworthy.
- : Bioassay Protocol Metadata Annotation: Proposed Standards Adoption, SLAS Discovery 29:100188 (2024) link
- : The Next Frontier of Environmental Unknowns: Substances of Unknown or Variable Composition, Complex Reaction Products, or Biological Materials (UVCBs), Environmental Science and Technology 56, 7448-7466 (2021) link
- : Using machine learning to parse chemical mixture descriptions, ACS Omega 6, 22400-22409 (2020) link
- : Capturing mixture composition: an open machine-readable format for representing mixed substances, Journal of Cheminformatics 11:33 (2019) link
- : Multiple Machine Learning Comparisons of HIV Cell-Based and Reverse Transcriptase Datasets, Molecular Pharmaceutics 11, 692-706 (2019) link
- : Ebola Virus Bayesian Machine Learning Models Enable New in Vitro Leads, ACS Omega 4, 2353-2361 (2019) link
- : Exploiting machine learning for end-to-end drug discovery and development, Nature Materials 18, 435-441 (2019) link
- : Assessment of Substrate Dependent Ligand Interactions at the Organic Cation Transporter OCT2 Using Six Model Substrates, Molecular Pharmacology 117 (2018) link
- : Accessible Machine Learning Approaches for Toxicology, Computational Toxicology: Risk Assessment for Chemicals 1-29 (2018) link
- : Developing Next Generation Tools for Computational Toxicology (Chapter 14), Computational Toxicology: Risk Assessment for Chemicals 363-387 (2018) link
- : Comparing Multiple Machine Learning Algorithms and Metrics for Estrogen Receptor Binding Prediction, Molecular Pharmaceutics 15, 4361-4370 (2018) link
- : Comparing and Validating Machine Learning Models for Mycobacterium tuberculosis Drug Discovery, Molecular Pharmaceutics 15, 4346-4360 (2018) link
- : Data Mining and Computational Modeling of High-Throughput Screening Datasets, Reporter Gene Assays 197-221 (2018) link
- : Small-molecule Bioactivity Databases, High Throughput Screening Methods (Chapter 16) 344-371 (2016) link
- : Collaborative drug discovery for More Medicines for Tuberculosis (MM4TB), Drug Discovery Today 22, 555-565 (2017) link
- : Machine Learning Model Analysis and Data Visualization with Small Molecules Tested in a Mouse Model of Mycobacterium tuberculosis Infection (2014-2015), Journal of Chemical Information and Modeling 56, 1332-1343 (2016) link
- : BioAssay templates for the semantic web, PeerJ Computer Science 2, e61 (2016) link
- : Open Source Bayesian Models. 3. Composite Models for Prediction of Binned Responses, Journal of Chemical Information and Modeling 56, 275-285 (2016) link
- : Making Transporter Models for Drug-Drug Interaction Prediction Mobile, Drug Metabolism and Disposition 43, 1642-1645 (2015) link
- : The Need for a Green Electronic Lab Notebook (Chapter 9), Green Chemistry Strategies for Drug Discovery 185-211 (2015) link
- : Open Source Bayesian Models. 2. Mining a 'Big Dataset' To Create and Validate Models with ChEMBL, Journal of Chemical Information and Modeling 55, 1246-1260 (2015) link
- : Open Source Bayesian Models. 1. Application to ADME/Tox and Drug Discovery Datasets, Journal of Chemical Information and Modeling 55, 1231-1245 (2015) link
- : Machines first, humans second: on the importance of algorithmic interpretation of open chemistry data, Journal of Cheminformatics 7, 9 (2015) link
- : Cheminformatics: Mobile Workflows and Data Sources (Chapter 14), The Future of the History of Chemical Information (ACS Symposium Series 1164) 237-253 (2015) link
- : The parallel worlds of public and commercial bioactive chemistry data, Journal of Medicinal Chemistry 58, 2068-2076 (2015) link
- : Hybrid machine-user learning system and process for identifying, accurately selecting and storing scientific data, US Patent #9594743 (2014) link
- : Bigger Data, Collaborative Tools and the Future of Predictive Drug Discovery, Journal of Computer Aided Molecular Design 28, 997-1008 (2014) link
- : Fast and accurate semantic annotation of bioassays exploiting a hybrid of machine learning and user confirmation, PeerJ 524 (2014) link
- : Mobile modeling in the molecular sciences, McGraw-Hill Yearbook of Science and Technology (2014) link
- : New target prediction and visualization tools incorporating open source molecular fingerprints for TB Mobile 2.0, Journal of Cheminformatics 6, 38 (2014) link
- : Looking Back to the Future: Predicting in Vivo Efficacy of Small Molecules versus Mycobacterium tuberculosis, Journal of Chemical Information and Modeling 54, 1070-1082 (2014) link
- : Rendering Molecular Sketches for Publication Quality Output, Molecular Informatics 32, 291-301 (2013) link
- : TB Mobile: a mobile app for anti-tuberculosis molecules with known targets, Journal of Cheminformatics 5, 13 (2013) link
- : Four disruptive strategies for removing drug discovery bottlenecks, Drug Discovery Today 18, 265-271 (2013) link
- : Cheminformatics workflows using mobile apps, Chem-Bio Informatics Journal 13, 1-18 (2013) link
- : Incorporating Green Chemistry Concepts into Mobile Chemistry Applications and Their Potential Uses, ACS Sustainable Chemistry and Engineering 1, 8-13 (2013) link
- : Open Drug Discovery Teams: A Chemistry Mobile App for Collaboration, Molecular Informatics 31, 585-597 (2012) link
- : Redefining Cheminformatics with Intuitive Collaborative Mobile Apps, Molecular Informatics 31, 569-584 (2012) link
- : Accurate Specification of Molecular Structures: The Case for Zero-Order Bonds and Explicit Hydrogen Counting, Journal of Chemical Information and Modeling 52, 3149-3157 (2011) link
- : Mobile apps for chemistry in the world of drug discovery, Drug Discovery Today 16, 928-939 (2011) link
- : Basic primitives for molecular diagram sketching, Journal of Cheminformatics 2, 8 (2010) link
- : 2D Depiction of Fragment Hierarchies, Journal of Chemical Information and Modeling, 50, 37-46 (2010) link
- : Detection and Assignment of Common Scaffolds in Project Databases of Lead Molecules, Journal of Medicinal Chemistry, 52, 469-483 (2009) link
- : 2D Depiction of Protein−Ligand Complexes, Journal of Chemical Information and Modeling, 47, 1933-1944 (2007) link
- : Flexible 3D pharmacophores as descriptors of dynamic biological space, Journal of Molecular Graphics and Modelling, 26, 622-633 (2007) link
- : 2D Structure Depiction, Journal of Chemical Information and Modeling, 46, 1107-1123 (2006) link
- : Bis(N,N'-dimethylimidazol-2-ylidene)mercury chlorotriiodomercury dimethyl sulfoxide solvate, Acta Crystallographica, C56, 26 (2000) link
- : Bromination and nitration reactions of metallated (Ru and Os) multiaromatic ligands and crystal structures of selected products, Journal of Organometallic Chemistry, 598, 262-275 (2000) link
- : Stepwise Conversion of an Osmium Trimethylstannyl Complex to a Triiodostannyl Complex and Nucleophilic Substitution Reactions at the Tin-iodine Bonds, Organometallics, 19, 1766-1774 (2000) link
- : The origin of the 'spike' in the EPR spectrum of C60-, Chemical Communications, 1229-1230 (2000) link
- : Cyclometallated complexes of ruthenium and osmium containing the o-C6H4PPh2 ligand, Journal of Organometallic Chemistry, 601, 299-304 (2000) link
- : Aryl Organometallic Compounds of Ruthenium and Osmium, Ph.D. Thesis (1999)
- : Electrophilic Aromatic Substitution Reactions at the Phenyl Ring of the Chelated 2-(2'-Pyridyl)phenyl) Ligand Bound to Ruthenium(II) or Osmium(II), Organometallics, 18, 2813-2820 (1999) link
- : 5-Bromination of an η2-8-Quinolyl Ligand Bound to Osmium(II) and Subsequent Lithiation and Derivatization of This Functionalized Ligand, Organometallics, 17, 4535-4537 (1998) link
- : Osmium complexes containing either chelating or non-chelating 8-quinolyl ligands, Journal of Organometallic Chemistry, 619, 545-546 (1997) link
- : Osmium nitrosyl complexes with osmium-tin bonds; Crystal structure of Os[Sn(p-tolyl)3](NO)(CO)2(PPh3), Journal of Organometallic Chemistry, 543, 543 (1997) link
- : Synthetic Approaches to Ruthenium and Osmium Stannyl Complexes, M.Sc. Thesis (1996)
Presentations and slides
2025
- Beyond Retrosynthesis: AI-powered reaction design: YouTube ⧉.
- Conjugate biomolecule datastructures for chemistry and biochemistry discovery: ACS Washington DC, Fall. Slides ⧉.
- Mixing small molecules and macromolecules in the world of informatics: ACS San Diego, Spring. Slides ⧉.
2024
- Some of the doors we unlock by describing molecules using generative models: ACS Denver, Fall. Slides ⧉.
- Assay annotation with ontologies; Introducing the DataFAIRy project: ACS Denver, Fall. Slides ⧉.
- Using ontologies to make bioassay protocols machine readable: ISMB conference, Montréal. Slides ⧉.
- Systematic annotation of bioassay protocols using ontologies: BioIT World, Boston. Slides ⧉.
2022
- Mixtures QSAR: modelling collections of chemicals: ACS San Diego, Spring. Slides ⧉.
2021
- Mixtures as first class citizens in the realm of informatics: Cambridge Cheminformatics. Slides ⧉.
- Mixtures InChI: A story of how standards drive upstream products: NIH Webinar. Slides ⧉.
2020
- Mixtures informatics for formulations and consumer products: RSC Webinar. Slides ⧉.
2019
- Capturing Mixtures — Bringing Informatics to the World of Practical Chemistry: CDD ⧉.
- Chemical mixtures: File format, open source tools, example data, and mixtures InChI derivative: ACS San Diego, Spring. Slides ⧉.
- Coordination InChI: Preliminary survey of inorganic compounds: ACS San Diego, Spring. Slides ⧉.
2018
- Bringing bioassay protocols to the world of informatics, using semantic annotations: ACS Boston, Fall. Slides ⧉.
- Leveling up chemical information for the era of big data: (CINF Luncheon Talk) ACS Boston, Fall. Slides ⧉.
2017
- Representing molecules with minimalism: A solution to the entropy of informatics: ACS Washington DC, Fall. Slides ⧉.
- Autonomous model building with a preponderance of well annotated assay protocols: ACS Washington DC, Fall. Slides ⧉.
- Expanding the target dimension: How to visualize a lot of SAR: ACS San Francisco, Spring. Slides ⧉.
2015
- Compact models for compact devices: Visualization of SAR using mobile apps: ACS Boston, Fall. Slides ⧉.
- The anatomy of a chemical reaction: Dissection by machine learning algorithms: ACS Boston, Fall. Slides ⧉.
2014
- Building a mobile reaction lab notebook: ACS Dallas, Spring. Slides ⧉.
- Cloud hosted APIs for cheminformatics designed for real time user interfaces: ACS Dallas, Spring. Slides ⧉.
- The Green Lab Notebook: Chemical intelligence and green informatics in a mobile app: ICCE, Toronto. Slides ⧉.
2013
- Putting together the pieces: building a reaction-centric electronic lab notebook for mobile devices: (Keynote) GDCh, Fulda. Slides ⧉.
- Cheminformatics workflows using the mobile + cloud platform: NETTAB, Venice. Slides ⧉.
- Mobile + cloud: a viable replacement for desktop cheminformatics?: ACS New Orleans, Spring. Slides ⧉.
- Pistoia Alliance App Strategy: Apps for life sciences R&D: ACS New Orleans, Spring. Slides ⧉.
- Open Drug Discovery Teams: NewTech meetup, Montréal. Slides ⧉.
2012
- Mobile chemistry apps participating in the open science revolution: ACS Philadelphia, Fall. Slides ⧉.
- Chemical structure diagrams, reactions and data: anytime, anywhere: ACS Philadelphia, Fall. Slides ⧉.
- Practical cheminformatics workflows with mobile apps: CINF Webinar. Slides ⧉.
- Building a mobile app ecosystem for chemistry collaboration: ACS San Diego, Spring. Slides ⧉.
- Open Drug Discovery Teams: Hacking Health, Montréal. Slides ⧉.
2011
- Chemistry in your pocket: ACS Anaheim, Spring. Slides ⧉.